Institute of Marine Biology, Biotechnology and Aquaculture - HCMR

RvLab
The R vLab makes use of “R” which is a statistical processing environment widely used by scientists working in many biodiversity related disciplines. It supports an integrated and optimized (in respect to computational speed-up and data manipulation) online R environment. This vLab tackles common problems faced by R users, such as severe computational power deficit. Many of the routines operating under the R environment, such as the calculation of several biodiversity indices and the running of the multivariate analyses, are often of high computational demand and cannot deliver a result when the respective datasets are in the form of large matrices.
Available after Sign In

MedOBIS vLab
MedOBIS is the Regional OBIS Node for the Mediterranean Sea. It is hosted by the Institute of Marine Biology, Biotechnology and Aquaculture https://imbbc.hcmr.gr/ (IMBBC), Hellenic Centre for Marine Research, HCMR https://www.hcmr.gr/en/, Heraklion (Crete). Launched in 2003, it has already been operational in 2005 as a Tier 3 Node of EurOBIS and covered the Eastern Mediterranean and the Black Sea. Under the European projects EMODNET and LifeWatchGreece (started in 2013), it became a Tier 2 node and extended to all Mediterranean Sea.
MedOBIS provides access to data from a wide range of sources and time periods, including new and historical data sets. MedOBIS actively contributes to global scientific efforts for FAIR and OPEN data.
The MedOBIS vLab consists of the MedOBIS IPT (Integrated Publishing Toolkit- http://ipt.medobis.eu/), which is available for sharing data and metadata, and Medobis viewer as a geodata tool, developed by open layers for visualization. MedOBIS can accept any data files from its data sources or data providers, and it publishes these data on its Integrated Publishing Toolkit (IPT), which is harvested by central OBIS. The Integrated Publishing Toolkit (IPT) is developed and maintained by the Global Biodiversity Information Facility (GBIF). For more information check here: https://obis.org/manual/contribute/
MedOBIS currently (2021) hosts 54 datasets, covering the period 1844 to 2017, with over 77,000 occurrence records accompanied with taxonomical, trait, geographical and environmental information. Login is required to access the vLab; while the IPT is open to any user.
Available without Sign In

Ecological Modeling
This vLab comprises of two online coupled models, which are parameterised and initialised for the specific conditions at a few specifically identified areas for which the required datasets exist. In an attempt to make the tool user friendly a graphic user interface (GUI) developed in the course of previous projects will be used. The GUI allows the user to view model results dynamically through any internet browser. Model results will be stored at the HCMR servers and the user will be able to select the area, scenario, and parameter required, which will then be returned as results in the form of plots. All model parameters and options will be available to the user online. The ultimate operation, therefore, of this vLab will be to allow the user to submit a request for the model to run under a different scenario than those already available.
Available without Sign In

Literature Mining
Biodiversity literature and data constitute a vast public resource open to mining and knowledge extraction. Associating organisms to key features of their life e.g. the environment in which they live, the way they feed, their breeding habits, is cornerstone in explaining biodiversity patters and informing ecological decisions. Eco-Systems Biology, and in particular network-based analysis, can provide holistic pictures of such associations, highlight novel relations and support hypothesis formulation and knowledge discovery. Initial aim of this vLab is to augment species related information based on data available in global biodiversity knowledge and literature aggregators, such as the Encyclopedia of Life (EOL) and the Biodiversity Heritage Library (BHL). Main focus of this virtual lab is the extraction of species - traits associations starting with the environment in which occur. Species and environments associations will be extracted by mining relevant text field clauses of: a) in-house LifewatchGreece data, b) the EOL and the BHL text collection. Also, interactive web-based visualizations will be developed to summarise the extracted species - environments association and support data exploration and landscape ecology studies.
Available without Sign In

Data Services
Data Services provide the users with tools in order to: a) publish their datasets and make them available to the community by providing information that allows a user to locate and access the resource and its curator/creator, b) search about datasets of interest by providing an efficient way of querying semantic networks and c) query and browse over the contents of the semantic network in an interactive manner, as well as using standard querying languages (i.e. SPARQL). The schema of the data that is provided by the users is mapped to the semantic model of the LWI and the data is transformed to LWI format before it is stored to the Infrastructure. The semantic model is based on CIDOC CRM (http://www.cidoc-crm.org/), CRM dig, CRM geo, CRM sci and MarineTLO (http://www.ics.forth.gr/isl/MarineTLO/).
Available without Sign In
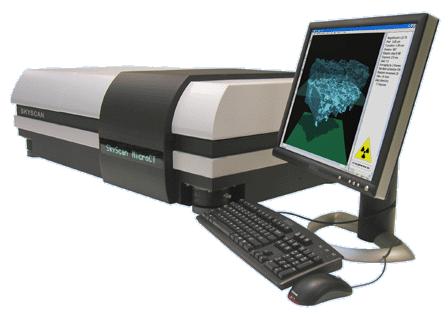
MicroCT Services
Micro-tomography (micro-computed tomography or microCT) is a method of non-destructive 3D x-ray microscopy, which allows the users to create 3D models of objects from a series of x-ray projection images, similar to the conventional clinical computer tomography. The MicroCT Service will offer a collection of virtual galleries of taxa which will be displayed and disseminated through a web-based framework, and will allow the user to manipulate the 3D models through a series of online tools or to download the datasets for local manipulations.
Available without Sign In

Genetic Services
The online Services of the Genetics working group are focused on 16S Metagenomics, which involve specific 16S ribosomal marker gene amplification by PCR. The 16S analysis is the most common approach for biodiversity studies. The Service focuses on taxonomic analysis from data derived from 454 Roche as well as Illumina sequencing technologies and provides the relevant software to allow analysis for both sequencing technologies. Efficient noise removal for 454 data is accomplished using the AmpliconNoise (http://qiime.org/scripts/ampliconnoise.html) pipeline and the taxonomic assignment for both de-noised 454 and Illumina data is achieved via QIIME (http://qiime.org/index.html).
Special Permission required after Sign In
Greek Taxon Information System (TIS) Services
This Service acts as the taxonomic backbone of the infrastructure: a taxonomic identity code (ID) is assigned to all the taxa registered in the GTIS. All the information (e.g. systematic classification, taxonomic description, functional trait information, geographical distribution, registered material in the systematic collections of the Museums, etc.), which is relevant to a taxon can be assigned to its ID and through this to any other kind of information linked to that. The GTIS Service is directly based on those developed in Flanders Marine Institute (VLIZ), which have been developed in the course of several EU and international projects and initiatives, such as MarBEF (Marine Biodiversity and Ecosystem Functioning), PESI (Pan-European Species directories Infrastructure) and WoRMS (World Register of Marine Species).
Available after Sign In

Biological Speciments Collection Services
There are many zoological, botanical, microbial, molecular and other types of collections spread in several academic and research establishments, such as zoological museums, botanical gardens, seed banks, etc. Many endemic terrestrial and marine species are included in these collections. Most of these collections have been digitised over the last years. This Service will make such collections available to the potential user from a single access point through the LifeWatchGreece site and will liaise them with similar collections from other EU countries.
Available after Sign In

Mobile Applications
Many of LifeWatchGreece web applications are also available as mobile applications. Currently there is a main application called "LifeWatch Greece" Mobile app that consists of 4 sub-applications, two of them available without login and the other two by using LifeWatchGreece portal credentials. Data from different Citizen Science projects such as Comber and CIGESMED can be seen in Citizen Science sub application. MicroCTvlab, RvLAB and MedOBIS are sub applications where the user has the ability to see most of the options that are available through the LifeWatchGreece portal. The "LifeWatch Greece" Mobile app is available for free at the Google Play Store.
Available without Sign In








